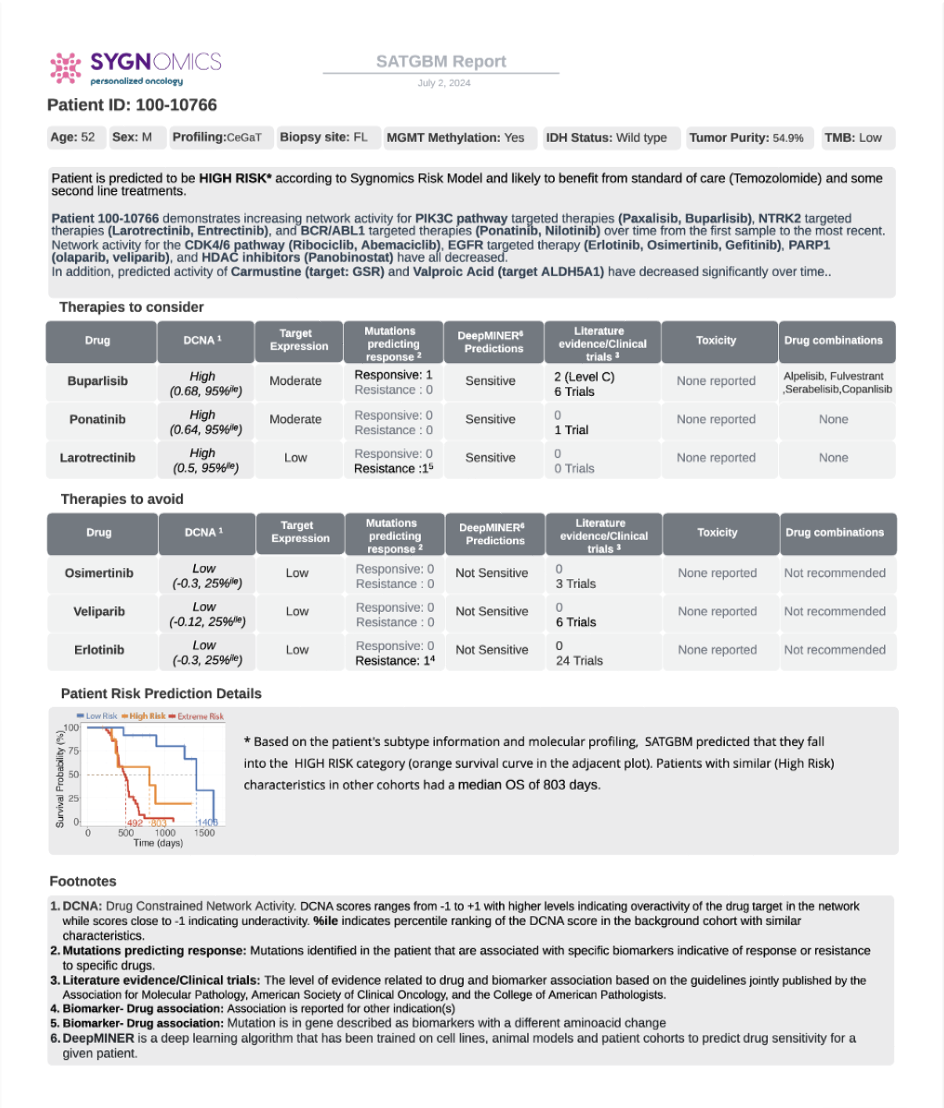
Example SAT report
Evidence of efficacy
Our systems approach significantly outperforms current clinical tests in risk stratification. In independent studies, cancer cells from both glioblastoma and multiple myeloma patients were significantly more sensitive to drugs recommended by our technology. And studies showed that glioblastoma patients stayed on their Sygnomics-recommended treatments up to five times longer than patients who received treatments deemed ineffective by our system.
It’s time to improve the statistics
SPOT-AI transforms cancer care from a population-level “drug-centric” approach to an N-1 “patient-centric” approach. While most cancer drugs fail in clinical trials and approved therapies work for fewer than 25% of patients, SPOT-AI reveals the true drivers of an individual’s cancer, helping oncologists make more effective, personalized treatment decisions and improving the patient’s outcomes.
Investigative Portals
Explore the portals that assisted in our research.

